Overview
Reproducible research is an important part of good scientific practice. Establishing robust computational analysis workflow facilitates reproducible research. There has been a strong emphasis on establishing graphic workflow management systems, such as Galaxy and Knime. However, the landscape and complexity of Linux command line tools for workflow management is vast and complicated. We aim to take advantage of European/International leaders in the field to present current scientific workflow paradigms and a lead workshops to implement a basic workflow. Topics covered include workflows and containerization.
Learning objectives
Docker
- Understand what containerisation is, and why you might use it in bioinformatics.
- Be familiar with Docker; basic concepts and structure.
- Find and run containers built by other people.
- Build your own application into a container (containerisation).
- Distribute your container online.
CWL
- Understand syntax and structure for CWL.
- Understand how to write CWL tool definitions for command line tools.
- Read and write CWL files written in YAML.
- How to run CWL workflows locally and on HYPATIA (formerly EG-CI).
- Join CWL tools into a workflow.
- Use Docker with CWL to provide software dependencies and ensure reproducibility.
Audience and requirements
This introductory tutorial is aimed towards bioinformaticians (graduate students and researchers), who are interested in becoming familiar to Docker based workflows in CWL, as is currently supported by the ΕLIXIR-Greece Compute Infrastructure (HYPATIA).
Prerequisites
- Experience in Shell; this includes basic commands (such as
ls,cp,mv,nano/vim) and operations such as (apt, installing tools etc).
Given that the workshop will offer hands-on training, participants will need to bring their own laptop with them.
SARS-CoV-2 guidelines
Please note that the venue requires that visitors follow the EU Digital COVID Certificate scheme as of the event date. Additional logistical details to help you plan your trip can be found on the event website. Please do not hesitate to get in touch with us if you need further assistance.
Maximum number of participants: 15 (will also have live streaming).
Registration
Date of the workshop: Monday, October 25th 2021
Please register to the workshop using this form.
Deadline: Wed 20/10/2021 12:00 local time
Notifications: Fri 22/10/2021
language of conduct: Greek
Instructors
- Kostis Zagganas IMSI, Athena Research Center
- Nikolaos Pechlivanis INAB, CERTH
- Thanasis Vergoulis IMSI, Athena Research Center
- Fotis Psomopoulos INAB, CERTH
Schedule
| Time | Details |
|---|---|
| 10:00 - 10:30 | Tutorial introduction |
| Part I: Containers Background | |
| 10:30 - 11:30 | Introduction to Containerization, run a container |
| Part II: Building a Container | |
| 11:45 - 13:15 | Building a container, distribute online |
| Part III: Basics of Workflow Languages - CWL | |
| 14:00 - 15:30 | Basics of workflow languages, writing CWL What is CWL? First example Running Tools Inside Docker |
| 15:30 - 16:45 | Combining tools in a workflow Writing workflows Scattering workflows |
| Part IV: Introduction to HYPATIA | |
| 17:00 - 18:15 | Using HYPATIA |
| 18:15 - 18:45 | Closing, discussion and Q&A |
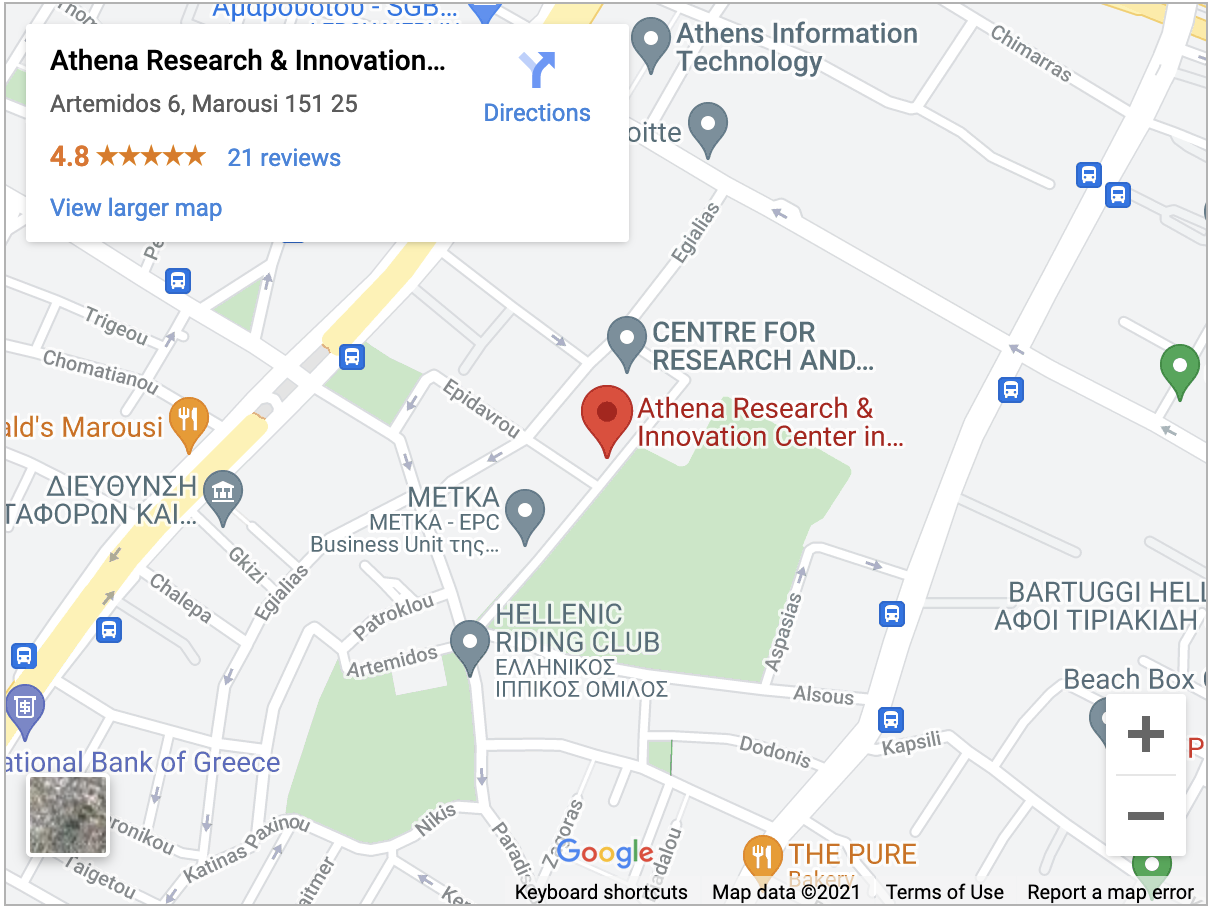
Venue
“ATHENA” Research Center (ATHENA RC)
Artemidos 6 & Epidavrou
15125, Marousi, GR
For more information, check the ATHENA RC contact page here.
License
This material is made available under the Creative Commons Attribution 4.0 International license. Please see LICENSE for more details.
Acknowledgements
We acknowledge support of this work by the project “ELIXIR-GR: Managing and Analysing Life Sciences Data” (MIS: 5002780) which is implemented under the Action “Reinforcement of the Research and Innovation Infrastructure”, funded by the Operational Programme “Competitiveness, Entrepreneurship and Innovation” (NSRF 2014-2020) and co-financed by Greece and the European Union (European Regional Development Fund).